Allowing foreigners and Israelis to fly to Israel from the US after the initial global outbreak of COVID-19 resulted in the spread of the virus throughout this country, according to Tel Aviv University researchers who mapped the spread by decoding the genomic sequence of the coronavirus strain in Israel.
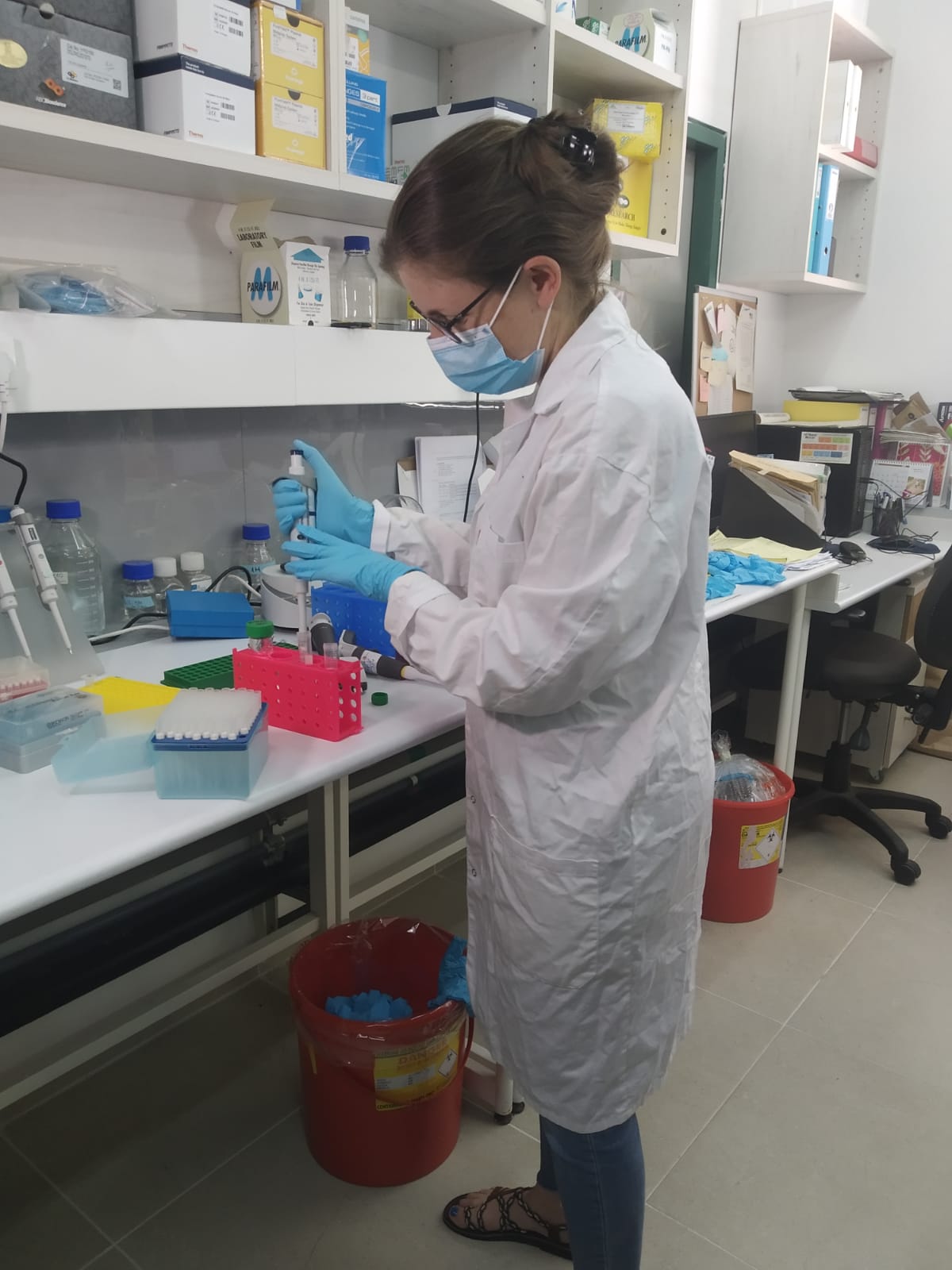
It was the first large-scale genomic sequencing of the novel coronavirus strain that has infected to date over 16,500 people in Israel.
Pressure from the US government on the Israeli government reportedly resulted in the delay in closing the gates to American passenger entry into Israel.
The virus did not come from China, and it barely originated in Europe and the Far East. Some 70% of the infections in Israel were caused by a SARS-CoV2 strain imported from the US and the remaining 30% by strains from: Belgium (8%), France (6%), the U.K. (5%), Spain (3%), Italy (2%), Australia (2%), the Philippines (2%) and Russia (2%).
The team, led by Dr. Adi Stern of the School of Molecular Cell Biology and Biotechnology at the university’s George Wise Faculty of Life Sciences, found that “super-spreaders” were responsible for most of the coronavirus cases in Israel.

“The novel coronavirus is characterized by mutations that occur at a set pace,” explained Stern. “These mutations do not affect the virus; it remains stable, but these mutations can help us trace the chain of infection from country to country. After the pandemic broke out in Wuhan, China, for example, one or two mutations occurred, and one virus with a mutation may have migrated to Europe where it experienced additional mutations, and from there it traveled to the US, and so on. We can look at these mutations as a kind of barcode that helps us keep track of the progression and transformation of the coronavirus as it moves from country to country.”
With policymakers in mind, the researchers developed a complex statistical model based on genomic sequencing that estimates the epidemiological parameters of viral spread.
The model shows that the rate of infection decreased significantly following strict quarantine measures taken in Israel and highlights a major discrepancy between the number of people each coronavirus patient infected. The model also estimates that over 80% of coronavirus cases in Israel were the direct result of only 10% of the coronavirus patients in Israel, meaning that these 10% were, in fact, super-spreaders. According to the model and to the genomic sequencing, Stern stated that no more than 1% of the population in Israel contracted the virus – a far cry from “herd immunity.”
The new genomic map provides insight into the precise spread of the novel coronavirus within Israel and will be useful in fighting any second wave of COVID-19. Until now, any assessment of the spread of infection relied on such subjective parameters as patient feedback. The new research will be able to expose the rate of infection in a household, in an apartment building, school, neighborhood and more. It will also provide early detection of super spreaders – people who travel far and wide and infect a large number of people – and could even identify major events with the potential to trigger widespread infection.
“Going forward,” noted Stern, the data obtained from genomic sequencing will serve as an important basis for informed decisions about which institutions to close, for what amount of time, and in which format. In our study, we performed the first massive genomic sequencing of the coronavirus in Israel,” she concludes. “This technology and the information it provides is of great importance for understanding the virus and its spread in the population, as a scientific and objective basis for local and national decision-making.”
The data obtained from the research can greatly help policymakers on issues such as closures and quarantines. In doing so, “the study makes a significant contribution to dealing with the epidemic in Israel, and, more importantly – we have developed tools that will allow us to cope, in real time, with the next outbreak that may occur,” the authors wrote.
To obtain a clear picture of the origin of infection in Israel, the researchers compared the genomic sequences of local patients to some 4,700 genomic sequences taken from patients around the world. They found that more than 70% of the patients had been infected by a coronavirus strain that originated in the US.
The scientists harnessed their genomic map to pinpoint mutations indicating where the virus originated from and later spread to within Israel. The study, published May 17 on medRxiv.org for immediate sharing with the global research community, is based on an analysis of the genomic sequences of over 200 patients at hospitals across Israel, who together constitute a representative sample of the general population.
The website, pronounced “med-archive,” is a free online archive and distribution server for complete but unpublished manuscripts in the medical, clinical, and related health sciences that have not been certified by peer review.
Tel Aviv University doctoral students Daniel Miller, Noam Harel, Talia Kostin, Omer Tirosh and Moran Meir conducted the research for the study in collaboration with scientists at Emory University, Gertner Institute, Sheba Medical Center at Tel Hashomer, the Holon Institute of Technology, Assuta Medical Center in Ashdod, Hadassah-University Medical Center in Jerusalem, Soroka University Medical Center in Beersheba, Barzilai Medical Center in Ashkelon, Poriya Medical Center in Tiberias, and the Genome Center at the Technion-Institute of Technology in Haifa.